TN, UNITED STATES, April 27, 2026 /EINPresswire.com/ — To overcome long-standing challenges posed by thick fungal cell walls, the high sample input demands of traditional ChIP-Seq, and excessive background noise, the team developed fCUT&Tag-Seq, a method that combines protoplast preparation with nuclear extraction to enhance antibody accessibility and introduces formaldehyde cross-linking to stabilize protein-DNA interactions. This approach enables high-resolution, high-sensitivity genome-wide mapping of histone modifications and chromatin-binding proteins across diverse fungi, providing a powerful tool for deciphering the epigenetic mechanisms underlying fungal gene regulation, development, and pathogenesis.
Fungal development, environmental adaptation, and pathogenesis all depend on the precise regulation of gene expression. Histone modifications and chromatin-binding proteins serve as key “switches” within this epigenetic regulatory network. However, genome-wide chromatin research in fungi, particularly in pathogenic filamentous species, has long been hampered by significant technical barriers. Traditional ChIP-Seq requires large amounts of starting material and is prone to chromatin epitope masking and high background noise. Meanwhile, the tough cell walls of filamentous fungi severely restrict antibody penetration and immunoprecipitation efficiency. These bottlenecks have constrained in-depth studies of fungal epigenetics and, more specifically, the regulatory mechanisms governing fungal pathogenicity.
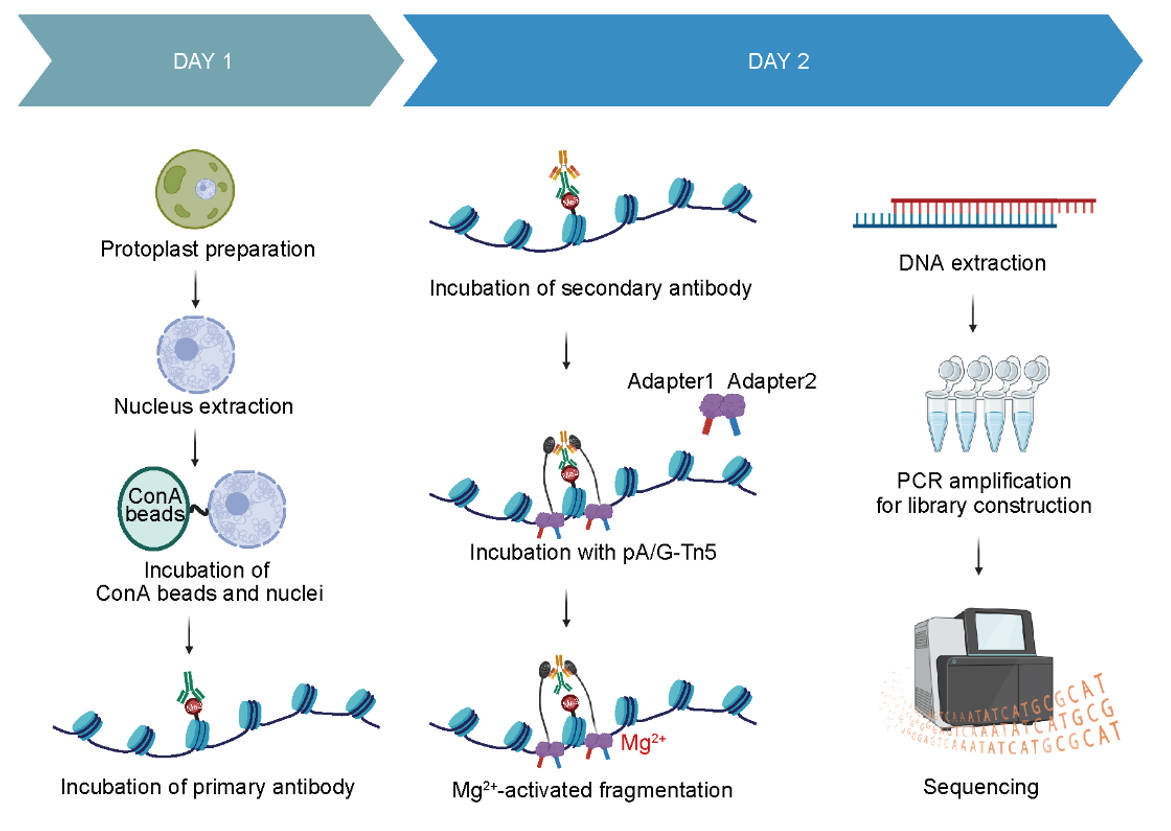
To address these challenges, the research team developed fCUT&Tag-Seq, a method specifically tailored for fungi. Building upon the conventional CUT&Tag protocol, this approach incorporates a series of systematic optimizations to accommodate the unique characteristics of fungal cells. First, the cell wall barrier is removed through enzymatic protoplast preparation, followed by gentle nuclear extraction to improve antibody access to target sites. Digitonin is then used to enhance nuclear membrane permeability, enabling efficient and precise target recognition and library construction while preserving the native chromatin state. For chromatin-binding proteins with lower abundance or weaker DNA-binding affinity, an additional formaldehyde cross-linking step is introduced to stabilize protein-DNA interactions and enhance detection sensitivity (Figure 1).
The study demonstrates that fCUT&Tag-Seq performs robustly across a range of filamentous and dimorphic fungi, including Verticillium dahliae, Neurospora crassa, Fusarium graminearum, and Sporisorium scitamineum. The method successfully mapped the genome-wide distribution of multiple histone modifications, including H3K9me3, H3K27me3, H3K4me3, and H3K18ac. It also enabled high-resolution profiling of chromatin-binding protein binding sites, including those of the histone acetyltransferase Gcn5 (Figure 2). Compared to traditional ChIP-Seq, fCUT&Tag-Seq exhibits a higher signal-to-noise ratio, superior reproducibility, and dramatically reduced sample requirements, needing as few as approximately 10,000 cells. The entire workflow can be completed in as little as two days.
This work provides a more efficient, sensitive, and accessible solution for fungal epigenomic research. Looking ahead, fCUT&Tag-Seq is expected to be extended to in planta (in situ) samples and unculturable fungi. This will empower researchers to uncover the dynamic changes in fungal chromatin, gene expression regulation, and pathogenesis-related molecular mechanisms under natural conditions, ultimately providing a theoretical foundation and potential targets for the prevention and control of major agricultural diseases.
Original Source URL
https://doi.org/10.1002/mlf2.70060
Funding information
This work was supported by the China NationalKey Research and Development Program (Grant No.2024YFC3406000), the Chinese Academy of Sciences (CAS)Projects for Young Scientists in Basic Research (Grant No.12YSBR‐080), the Strategic Priority Research Program ofChinese Academy of Sciences (Grant No. XDA0450000) toC‐M.S., and the Young Scientists Fund of the NationalNatural Science Foundation of China (Grant No. 32402338) to H.W.
Lucy Wang
BioDesign Research
email us here
Legal Disclaimer:
EIN Presswire provides this news content “as is” without warranty of any kind. We do not accept any responsibility or liability
for the accuracy, content, images, videos, licenses, completeness, legality, or reliability of the information contained in this
article. If you have any complaints or copyright issues related to this article, kindly contact the author above.
![]()
Media gallery